Attention has been increasing focused on the role of the bone marrow microenvironment in the pathogenesis and progression of hematological malignancies, including myelodysplastic syndromes (MDS) and acute myeloid leukemia (AML).1–3 Thus, the identification of genes or proteins that are differentially expressed in the abnormal bone marrow niche may provide new therapeutic opportunities and perspectives on the biology of these diseases. In a recent report by Fei et al.,4 bone marrow mesenchymal stromal cells (BMMSCs) from MDS patients presented an upregulation of p21 and p53 expressions, suggesting that activation of this pathway may contribute to the senescent behavior of these cells.4 In contrast, another study that also analyzed the expression profile of senescence-related genes in BMMSCs from MDS and healthy donors observed a downregulation of CDKN1A expression, but no modulation in TP53 expression.5
To provide additional evidence on p21 and p53 expression in MDS BMMSCs, we verified the expression of these genes in a cohort of eight healthy donors with median age of 45 years (range: 28–57), 23 MDS patients with median age of 70 years (range: 16–90), seven AML patients with myelodysplasia-related changes (AML-MRC) with median age of 69 years (range: 30–86) and 12 de novo AML cases according to the WHO 2008 classification, with median age of 61 years (range: 44–82).6 The MDS group was comprised of one refractory cytopenia with unilineage dysplasia (RCUD), four refractory anemia with ringed sideroblasts (RARS), 12 refractory cytopenia with multilineage dysplasia (RCMD), two refractory anemia with excess blast-1 (RAEB-1) and four refractory anemia with excess blast-2 (RAEB-2). Bone marrow mononuclear cells were isolated by Ficoll-Hypaque plus density-gradient centrifugation (GE Healthcare, Uppsala, Sweden) and cultured in Dulbecco's Modified Eagle's Medium (DMEM; Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS), penicillin/streptomycin and maintained at 37°C, normoxia with 5% CO2. After the fourth passage, all patient and control-derived BMMSCs presented a homogeneous cell population (negative for CD31, CD34, CD45 and HLA-DR, and positive for CD73, CD90 and CD105), confirming their mesenchymal origin, according to the International Society for Cellular Therapy.7 Next, these samples were used for gene expression analysis by quantitative polymerase chain reaction (qPCR) in a ABI 7500 Sequence Detection System (Applied Biosystem, Foster City, CA, USA) using specific primers for CDKN1A (p21 – FW: TGTCACTGTCTTGTACCCTTGT; RV: GCCGGCGTTTGGAGTGGTAG), TP53 (p53 – FW: GGCGCACAGAGGAAGAGAAT; RV: GGAGAGGAGCTGGTGTTGTTG), and HPRT1 (FW: GAACGTCTTGCTCGAGATGTGA; RV: TCCAGCAGGTCAGCAAAGAAT). The relative gene expression was calculated using the equation, 2−ΔΔCT.8 Statistical analyses were performed by ANOVA and Bonferroni post-test using GraphPad Prism 5 (GraphPad Software, Inc., San Diego, CA, USA); a p-value <0.05 was considered statistically significant.
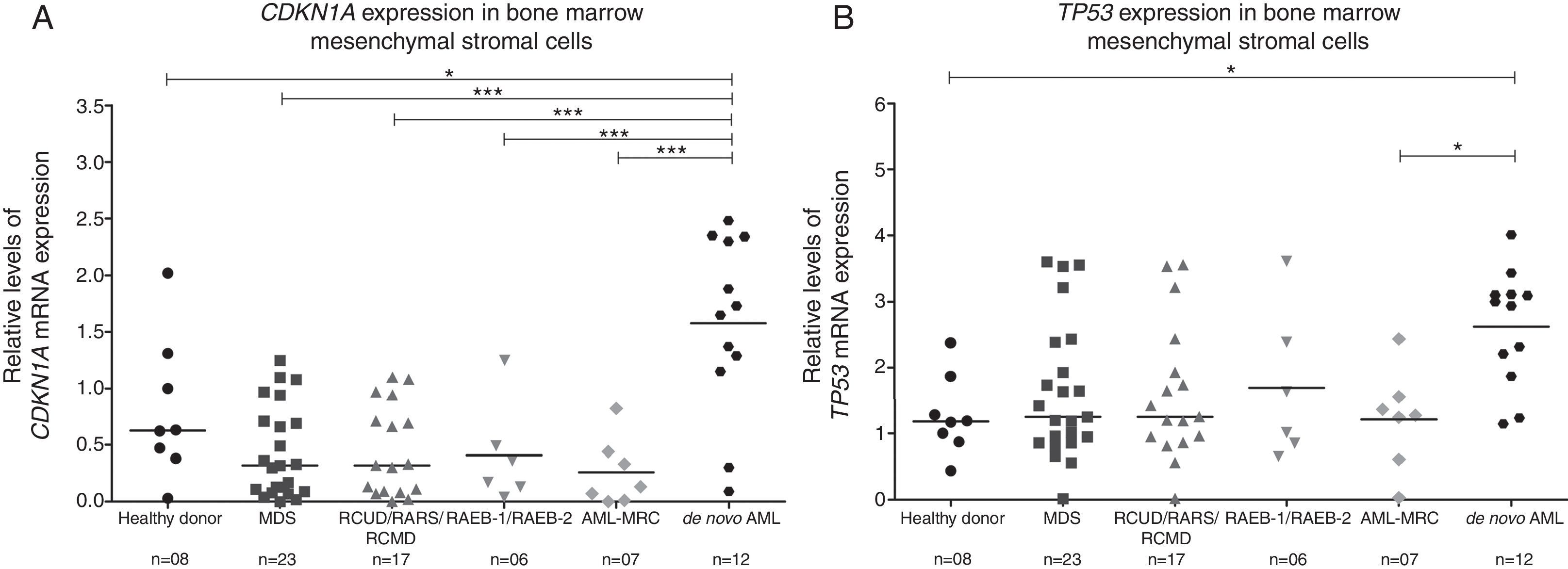
In our cohort, BMMSCs from de novo AML presented increased CDKN1A mRNA levels compared to healthy donors, MDS and AML-MRC patients (p-value <0.05; Figure 1A). TP53 expression was also higher in de novo AML compared to healthy donors and AML-MRC patients (p-value <0.05; Figure 1B). However, we observed no differences in CDKN1A and TP53 mRNA levels in BMMSCs from MDS and AML-MRC patients compared to healthy donors (p-value >0.05; Figure 1). When MDS patients were stratified by the WHO 2008 classification, no differences were observed between RCUD/RARS/RCMD, RAEB-1/RAEB-2, AML-MRC and healthy donors (p-value >0.05; Figure 1).
CDKN1A and TP53 expression in bone marrow mesenchymal stromal cells from myelodysplastic syndromes, acute myeloid leukemia with myelodysplasia-related changes, de novo acute myeloid leukemia and healthy donors.
Quantitative polymerase chain reaction analysis of CDKN1A (A) and TP53 (B) mRNA expression in bone marrow mesenchymal cells. The HPRT1 gene was used as an endogenous control and a healthy donor was used as a calibrator sample. Horizontal lines indicate medians. *p-value <0.05, **p-value <0.01, ***p-value <0.001; ANOVA test and Bonferroni post-test.
Existing data on MDS-derived BMMSC biology are somewhat controversial, reflecting differences in methodologies and the heterogeneity of patients.9 There is evidence that different culture media can significantly influence the phenotype of mesenchymal cells.10,11 Thus, the lack of optimization of cell isolation techniques and expansion conditions may influence the gene expression profile and the identification of specific markers in BMMSC studies. In the protocol used by Fei et al.,4 BMMSCs were cultured in a special human mesenchymal stem cell growth medium, while Pavlaki et al.5 cultured BMMSCs using DMEM.
With regard to the CDKN1A and TP53 expression profile in de novo AML BMMSCs, our results corroborate the findings of Ruvolo et al.,12 who observed an upregulation of p21 and p53 in BMMSCs from AML, suggesting increased cellular senescence. Importantly, our results highlight the biological and molecular differences between AML-MRC and de novo AML reported by other research groups.13–15
Several lines of evidence indicate that alterations in the bone marrow niche contribute to the development and progression of MDS and AML and one of the most characterized elements of the bone marrow niche is the BMMSCs.9,16 This cell population represents a small fraction of bone marrow nucleated cells, and a standardized protocol for isolation and culture of BMMSCs may be necessary to minimize experimental variations and provide conclusive information about the biology of these cells in these hematological malignancies. Our results suggest that upregulation of CDKN1A and TP53 may indicate senescence of de novo AML BMMSCs, which may contribute to the ineffective hematopoiesis found the this disease.
Conflicts of interestThe authors declare no conflicts of interest.
Funding for this work was provided by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP). The authors would like to thank Dr. Nicola Conran for the English review.