Nucleotide excision repair pathway (NER) is an essential mechanism for single-strand breaks (SSB) repair while xeroderma pigmentosum family (XPA to XPG) is the most important system to NER. Myelodysplastic syndrome (MDS) is a heterogeneous hematological cancer characterized by cytopenias and risk of acute myeloid leukemia (AML) transformation. MDS pathogenesis has been associated with problems of DNA repair system. This report aimed to evaluate NER polymorphisms (XPA rs1800975, XPC rs2228000, XPD rs1799793 and XPF rs1800067) in 269 MDS patients of different populations in Latin America (173 Brazilian and 96 Argentinean). Genotypes were identified in DNA samples by RT-qPCR using TaqMan SNP Genotyping Assay. Regarding rs1799793 polymorphism of XPD for Brazilian population, the heterozygous genotype AG presented a high odds ratio (OR) to have a normal karyotype (p = 0.012, OR=3.000) and the mutant homozygous genotype AA was associated to a high OR of AML transformation (p = 0.034, OR=7.4). In Argentine population, the homozygous mutant AA genotype of rs1800975 polymorphism of XPA was associated with an increased odd to have hemoglobin levels below 8g/dL (p = 0.013, OR=10.000) while for the rs1799793 polymorphism of XPD, the heterozygous AG genotype decreased OR to be classified as good (p < 0.001, OR=9.05 × 10−10), and intermediate (p < 0.001, OR=3.08 × 10−10), according to Revised-International Prognostic Scoring System. Regarding the rs1800067 polymorphisms of XPF, the homozygous mutant AA genotype showed a decreased OR to be classified as good (p < 0.001, OR=4.03 × 10−13) and intermediate (p < 0.001, OR=2.54 × 10−13). Our report reinforces the heterogeneity of MDS and demonstrates the importance of ethnic differences and regional influences in pathogenesis and prognosis of MDS.
Cells are continuously susceptible to DNA damage caused by endogenous and exogenous factors, which may lead to cancer development if not properly repaired. Different repair mechanisms are involved in DNA damage response, such as nucleotide excision repair (NER).1–4 NER pathway is an essential mechanism for single-strand breaks (SSB) recognizing large amount of DNA single-strand breaks and crucial in maintaining genomic stability. Xeroderma pigmentosum family (XPA to XPG) are the most important genes related to NER repair, responsible for promoting and coordinating it.3,4
Functional polymorphisms (FPs) of DNA repair system are likely to modify the risk of DNA damage and cancer development5,6 and studies have indicated associations of FPs with differences in susceptibility and prognosis of Myelodysplastic syndrome (MDS).7–11 We previously demonstrated the importance of ATM polymorphism12 and new functional polymorphisms of the Xeroderma Pigmentosum family (XPA rs1800975, XPC rs2228000 and XPD rs1799793) in MDS pathogenesis.13,14
MDS is a very heterogeneous bone marrow cancer characterized by bone marrow insufficiency, which leads to ineffective hematopoiesis with cytopenias, bone marrow dysplasias and an increased risk of progression to acute myeloid leukemia (AML).15,16 The pathogenesis has been linked to DNA damage in hematopoietic stem cells and it is highly related to genotoxic factors.17,18
In 2005, Matsuda et al.,19 evaluated differences in clinical features between Asian and Western MDS cases. They demonstrated Japanese patients were younger than German patients, had more severe cytopenias and lower cumulative risk of AML evolution than German patients. Since them, a lot of reports have demonstrated how heterogeneous MDS can be according to people evaluated. In 2015, we reported the first study from South-America, which described clinical features, demographic, overall survival and AML transformation of 1080 MDS patients from Argentina (Ar), Brazil (Br) and Chile (Ch). We demonstrated Brazilian series had higher predominance of Refractory Anemia Ring Sideroblasts (RARS) subtype than Ar and Ch, hemoglobin levels were lower in Brazilian and Chilean series than Ar and overall survival was 35, 56 and 55 months for Chile, Argentine and Brazil respectively.20
Due to this very impressive regional difference in MDS which contributes to its high heterogeneity, we decided to evaluate functional polymorphisms of genes related to genomic stability, more precisely: components of NER (XPA rs1800975, XPC rs2228000, XPD rs1799793 and XPF rs1800067 polymorphisms). We believe these analysis help understand the epidemiological characteristics which make MDS so heterogeneous.
Patients, materials and methodsPatientsWe evaluated 269 MDS patients from Latina America (173 from Brazil, 96 from Argentina) diagnosed according to the World Health Organization (WHO)21 and the Revised International Prognostic Scoring System (IPSS-R).22 Two hundred fifty-four (254) controls, sex and age-matched, were used (154 from Brazil, 100 from Argentina) for analysis. Among Brazilian patients, 99 were from Northeast and 78, Southeast.
We evaluated associations between functional polymorphisms of XPA rs1800975, XPC rs2228000, XPD rs1799793 and XPF rs1800067 with variables gender, age, cytopenias (number of cytopenias, hemoglobin level, neutrophil and platelet count), blast percentage, karyotype, bone marrow cellularity, ring sideroblast percentage, transfusional dependency, WHO 2016 classification21 IPSS-R score,22 and AML evolution. This study was approved by the research ethics committee CAAE #03258718.3.0000.5054.
Cytogenetic analysisFor the conventional G-Banded karyotype analysis, cultures were established in RPMI 1640 medium (Gibco, Grand Island, NY, USA) containing 30% fetal calf serum. After a 24 h incubation, colcemid was added to cell culture for blocking of mitotic fuse (final concentration 0.05 μg/mL). After harvesting procedure, the cells were exposed to KCl hypotonic solution (0.068 mol/L) and fixed with Carnoy's buffer (methanol/acetic acid in 3:1 proportion). The slides were prepared and stained with Giemsa solution. A minimum of 20 metaphases were analyzed whenever possible using CytoVision Automated Karyotyping System (Applied Imaging, San Jose, CA, USA).
DNA extractionGenomic DNA was extracted using TRIzol Reagent™ (Invitrogen, Carlsbad, CA, USA), according to the manufacturer's protocol.
Allelic discrimination by Real Time-PCR (RT-PCR)Genotypes were identified in DNA samples by real-time polymerase chain reaction (RT-PCR), using TaqMan SNP Genotyping Assay (ThermoFisher Scientific, Foster City, CA, USA). We performed an allelic discrimination assay for rs1800975 (C_482935_1_), rs2228000 (C_16018061_10), rs1799793 (C_3145050_10) and rs1800067 (C_3285104_10) polymorphisms using the TaqMan Genotyping Master Mix kit® (catalog #4371355, ThermoFisher Scientific, Foster City, CA, USA), according to the manufacturer's protocol.
Each 10μL PCR reaction contained ∼30ng of genomic DNA in final volume of 4.5 µL, 5.0 µL of 2X TaqMan Genotyping Master Mix and 0.5 µL of 20X TaqMan Genotyping assay mix. The following thermal cycling conditions were: an initial activation at 95 °C for 10 min., followed by 40 cycles of 95 °C for 15 s of denaturation step at 95 °C and annealing/enzyme extension at 60 °C for 1 min. PCR was performed using 7500 Fast System® (Applied Biosystems, Carlsbad, CA, USA).
Statistical analysisPearson chi-square test (x2) was used to determine the Hardy–Weinberg (H-W) equilibrium and the differences in genotypes distribution. All genotypes were divided and analyzed in three distinct genetic models, according to the study of Clarke et al.23: 1. Genotype distribution model (wild-type versus heterozygous versus mutated); 2. Dominant genetic model (wild-type versus heterozygous + mutated) and 3. Recessive genetic model (mutated versus wild-type versus heterozygous + wild-type). Differences in allele and genotype frequencies and comparisons of variables between the control and MDS groups were evaluated using the Pearson's chi-square test or Fisher's exact test. Multinomial logistic regression analysis was used to measure the association between an exposure and an outcome (odds ratio-OR) with 95% confidence intervals (CI). Statistical analysis was performed using the SPSS 20.0 (SPSS Inc., Chicago, IL, USA).
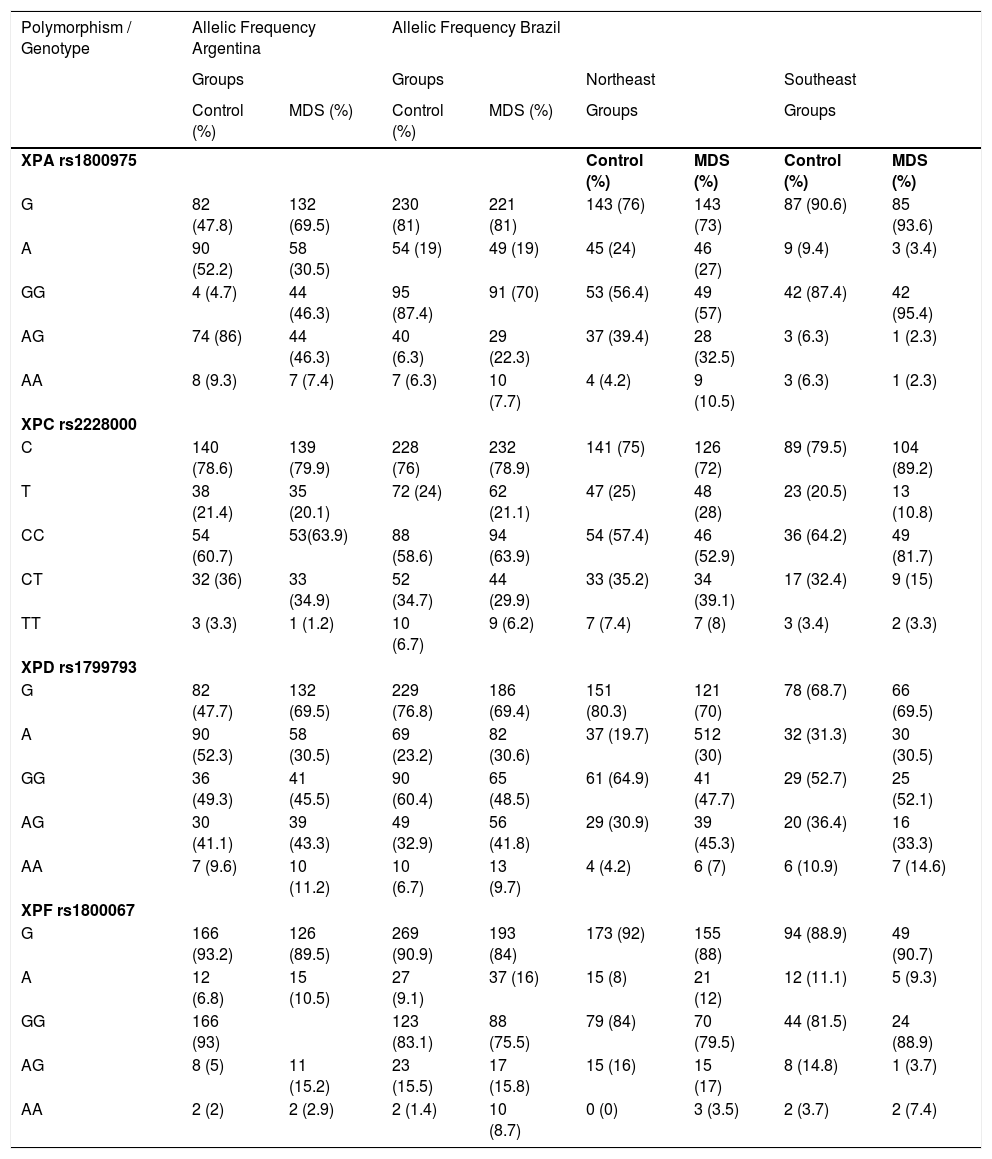
ResultsGenotype frequencies and allele distributions of NER polymorphismsThe distributions of allele and genotype frequencies of XPA rs1800975, XPC rs2228000, XPD rs1799793 and XPF rs1800067 of 269 MDS patients (173 from Brazil and 96 from Argentina) and 254 healthy volunteers (154 from Brazil and 100 from Argentina) are presented in Table 1.
Distribution of genotypes in MDS patients and healthy controls according to the allelic frequency.
G and C: wild allele; A and T: mutant allele; GG and CC: wild homozygote; AG and CT: heterozygote; AA and TT: mutant homozygote
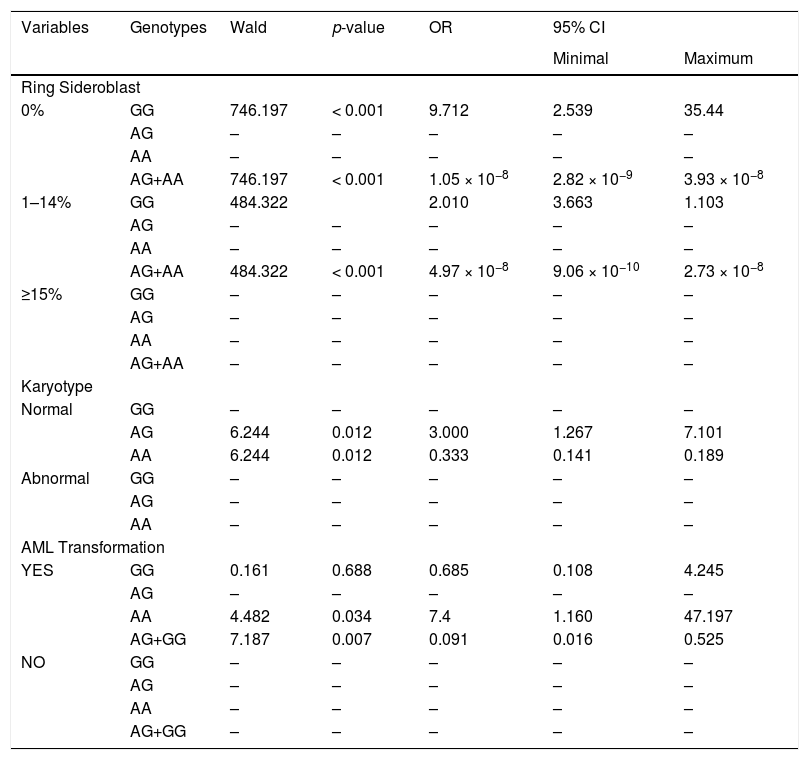
For the rs1799793 polymorphism of XPD, the wild homozygous GG genotype was associated with a decreased OR of being classified as MDS with ring sideroblasts according to WHO 2016 (p = 0.050, OR=0.91, CI 0.008-0.995). The presence of mutant allele A (AG+AA) was associated with a higher chance for MDS with ring sideroblasts (p = 0.050, OR=11.00, CI=1.005-120.430) (Table 2).
Multinomial Logistic Regression of XPD rs1799793 SNP genotypes in Brazil.
GG: wild homozygote, AG: heterozygote and AA: mutant homozygote.
Regarding karyotype, the heterozygous genotype AG presented a higher odds ratio to have a normal karyotype (p = 0.012, OR=3.000, CI=1.26-7.1) than mutant genotype AA showed (p = 0.012, OR=0.333, CI=0.141–0.189) (Table 2). Regarding AML transformation, we found a significant association between the mutant homozygous genotype AA and a higher odds ratio of AML transformation (p = 0.034, OR=7.4, IC=1.160–147). The recessive model combination AG+GG was associated with a lower odds ratio of AML transformation (p = 0.007, OR=0.091, CI=0.016-0.525) (Table 2).
We found no significant associations between the XPD rs1799793 polymorphism and the variables age, cytopenias, bone marrow cellularity, blast percentage, IPSS-R prognosis,22 transfusional dependency, and death (p>0.05).
In a second step, we evaluated the rs1799793 polymorphism of XPD in Brazilian regions separately (Northeast and Southeast). Regarding Northeast population, we found an association between the wild genotype GG with a decreased odds ratio of developing MDS in the dominance model (p = 0.009, OR=0.445, CI=0.242-0.816). Regarding Southeast population, we found a significant association between the heterozygous AG genotype with a higher odds ratio to have normal karyotype (p = 0.018, OR=7.583, CI=1.41–40.54) than mutant AA genotype (p = 0.930, OR=1.083, CI=0.182–6.439).
XPA rs1800975 polymorphismsThe genotypes analyzed for rs1800975 polymorphisms were not in Hardy-Weinberg equilibrium (p> 0.05) in Brazilian population, precluding multinomial logistic regression.
XPC rs2228000 polymorphismThe genotypes analyzed for rs2228000 polymorphisms were not in Hardy-Weinberg equilibrium (p > 0.05) in Brazilian population, precluding multinomial logistic regression.
XPF rs1800067 polymorphismsThe genotypes analyzed for rs1800067 polymorphisms were not in Hardy-Weinberg equilibrium (p > 0.05) in Brazilian population, precluding multinomial logistic regression.
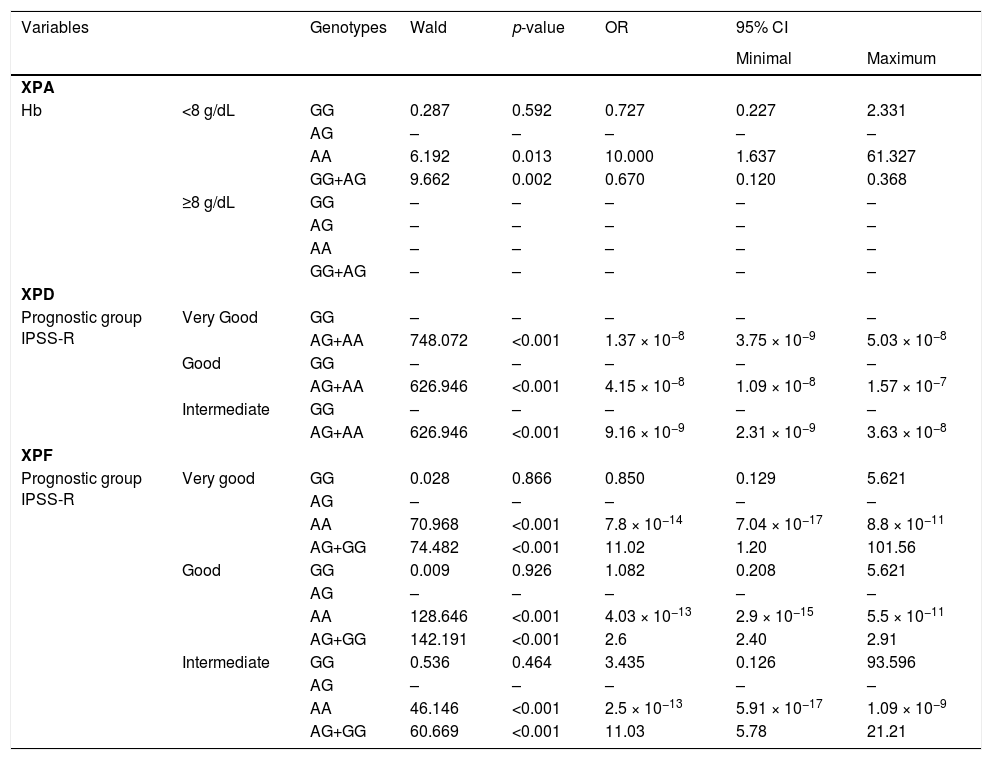
Analysis of NER polymorphisms – ArgentinaXPA rs1800975 polymorphismsRegarding XPA rs1800975 polymorphism, the homozygous mutant AA genotype was associated with an increased odd to have hemoglobin levels below 8g/dL (p = 0.013, OR=10.000, IC=1.63–61.32). As expected, the GG+AG combination of recessivity model was associated with a lower odds ratio of Hb <8g/dL (p = 0.002, OR=0.67, CI=0.12–0.368) (Table 3).
Multinomial logistic regression of SNP in Argentina.
GG: wild homozygote, AG: heterozygote and AA: mutant homozygote.
We found no significant associations between the XPA rs1800975 polymorphisms and variables of gender, age, cytopenias (number of cytopenias, neutrophil and platelet count), blast percentage, karyotype, bone marrow cellularity, ring sideroblast percentage, transfusional dependency, WHO 2016 classification21 IPSS-R score,22 and AML evolution (p > 0.05).
XPD rs1799793 polymorphismFor the rs1799793 polymorphism of XPD, we found a significant association between the heterozygous AG genotype and the decreased odds ratio to be classified as very good (p < 0.001, OR=4.7 × 10−10, CI=1.95 × 10−11-1.13 × 10−8), good (p < 0.001, OR=9.05 × 10−10, CI=4.76 × 10−11-1.72 × 10−18), and intermediate (p < 0.001, OR=3.08 × 10−10, OR=1.26 × 10−11-7.54 × 10−9), according to the Revised International Prognostic Score System.22 The interaction of AG+AA genotypes were also associated with decreased odds ratio to be classified as very good (p <0.001, OR=1.37 × 10−8, CI=3.75 × 10−9-5,03 × 10−8), good (p < 0.001, OR=4.15 × 10−8, CI=1.09 × 10−8-1.57 × 10−7) and intermediate (p < 0.001, OR=9, 16 × 10−9, CI=2.31 × 10−9-3.63 × 10−8) according to IPSS-R score (Table 3).
We found no significant associations between the XPD rs1799793 polymorphism and variables gender, age, cytopenias (number of cytopenias, hemoglobin, neutrophil and platelet count), blast percentage, karyotype, bone marrow cellularity, ring sideroblast percentage, transfusional dependency, WHO 2016 classification21 and AML evolution (p>0.05).
XPF rs1800067 polymorphismsRegarding the rs1800067 polymorphisms of XPF, the homozygous mutant AA genotype showed a decreased odds ratio to be classified as very good (p < 0.001, OR=7.88 × 10−14, CI=7.04 × 10−17-8.81 × 10−11), good (p < 0.001, OR=4.03 × 10−13, CI=2.908 × 10−15-5.58 × 10−11) and intermediate (p < 0.001, OR=2.54 × 10−13, CI=5.91 × 10−17-1.09 × 10−19) according to IPSS-R. (Table 3). In the recessive model, there was significance in the interaction of AG+GG genotypes with a higher odds ratio of very good (p < 0.001, OR=11.02, CI=1.20-101.564), good (p < 0.001, OR=2.6, CI=2.40-2.91) and intermediate (p <0.001, OR 11.07, CI 5.78 - 21.21) group of IPSS-R22 (Table 3).
We found no significant associations between the XPF rs1800067 polymorphism, and variables of gender, age, cytopenias (number of cytopenias, hemoglobin, neutrophil and platelet count), blast percentage, karyotype, bone marrow cellularity, ring sideroblast percentage, transfusional dependency, WHO 2016 classification,21 and AML evolution (p>0.05).
XPC rs2228000 polymorphismNo association was detected for this polymorphism in cases from Argentina.
DiscussionWe evaluated single nucleotide functional polymorphisms (polymorphisms that have been reported to influence gene functions) of NER pathway which occur with high frequency and have been previously associated with cancer development: XPA rs1800975, XPC rs2228000, XPD rs1799793 and XPF rs1800067.14
Brazilian patients are from distinct regions of Brazil, the Northeast and Southeast. We found relevant associations between the variants rs1799793 polymorphism of XPD, which was the only one in Hardy-Weinberg equilibrium in patients from Brazil. The presence of adenine as polymorphic variant in mutant phenotype (AG or AA) increased chance of being classified as MDS with ring sideroblasts (WHO 2016 21). This subtype is considered a low-risk disease with very distinct pathogenesis characterized by splicing problems, especially related to the presence of SF3B1 mutations.21 In a multicenter retrospective analysis of 1080 patients with de novo MDS from Brazil (Br), Argentina (Ar) and Chile (Ch),20 we observed higher frequency of MDS with ring sideroblasts in Brazilian patients than Argentina and Chile. Br series showed 18% vs 10% Ar vs 1% Ch (p < 0.001) of MDS with ring sideroblasts and 50% of these patients were farmers from the rural area of Brazil. This region is less favored economically and these farmers do not have access to personal protective equipments (PPE) what increases the level of exposure to pesticides and other toxic environmental factors, commonly associated with the etiology of MDS, especially the subtype of MDS with ring sideroblasts.20 Although we are not able to explain the pathogenesis linking this mutant phenotype of XPD to increased chance of being MDS with ring sideroblasts, we suggest this mutant allele decreases the activity of XPD, possibly creating genomic instability. XPD helicase is an essential component of NER repair pathway of single-stranded lesion and its function is critical for the correction of damage caused by tobacco and other carcinogens (probably pesticides).18,20
In fact, we also detected the mutant AA (rs1799793) was associated with increased odds of AML transformation and a decreased odds ratio of normal karyotype. The presence of the wild variant (GG) may have a protective effect on genomic stability of these patients. Thus, we suggest presence of AA polymorphic homozygous genotype for the XPD gene may be a candidate of poor prognosis in MDS patients from Brazil.
XPA, XPC and XPF genotypes were not in Hardy-Weinberg equilibrium which precluded its evaluation for MDS from Brazil. The Hardy-Weinberg theorem states that allele and genotype frequencies in a population will remain constant from generation to generation in the absence of other evolutionary influences. Gene frequencies may be due, principally, to systematic processes such as gene flow, natural selection and mutations.23 Brazil presents continental dimension which created different development and colonization process for each region, generating a highly ethnically mixed population. We know Brazil ethnic composition is derived from three major ethnic groups: Indigenous, African, and European. According to a meta-analysis by Pena et al.,24 the South and Southeast regions have 77% of European genomic ancestry, while the Northeast, 60.6%. The opposite can be seen when looking at African contribution: 12.7% in the South, 18.9% in Southeast and 30.3% in the Northeast. Probably, all these facts contributed to absence of Hardy-Weinberg equilibrium here detected.
We found interesting associations between the genotypes of the XPA, XPD, and XPF and characteristics of patients from Argentina. The presence of the homozygous AA polymorphic variant of XPA is associated with an increased risk of severe anemia (hemoglobin <8g/dL). More than two-thirds of patients present anemia at diagnosis with a hemoglobin level <10g/dL and about 85% of the patients develop more severe anemia during follow up.21 Patients with hemoglobin <8g/dL are considered red blood cell transfusion-dependent and severe anemia increases the harmful effects of comorbidities.22 The main cause of death among MDS patients not RAEB is cardiac dysfunction. We previously demonstrated cardiac dysfunction as a marker of grim prognosis in transfusion dependent MDS patients.25 Using tissue doppler echocardiography, we detected preclinical markers of cardiac lesion in MDS with average values of ventricular end-systolic and end-diastolic volumes in transfusion dependent MDS group higher than not dependent and controls and theses abnormalities were strongly inversely correlated to hemoglobin levels.22
We found similar results for XPD and XPF polymorphisms. The presence of mutant allele A, even if presented as heterozygous genotype, was associated with a decreased chance of the patient being classified as very good, good or intermediate prognosis according to IPSS-R. Therefore, for both XPA, XPD, and XPF genes, the presence of polymorphic variants was related to, according to IPSS-R,22 lower overall survival and increased chance of AML transformation. The presence of these polymorphic variants (XPA, XPD and XPF genes in Argentina) affects the control of the genomic stability, which may be associated with a more aggressive disease. A higher frequency of mutant A allele in XPA gene was observed in Argentina (30.5%) compared to Brazil (19%), which may justify the lower percentage of Argentinean patients in the lower-risk groups.
MDS is distinct in different continents and populations. Miyazaki et al. (2018) 26 analyzed an extensive international database, collected by the International Working Group for Prognosis of MDS, and showed differences between Japanese and Caucasians (American or European). Asian patients were significantly younger, with more severe cytopenias and higher IPSS-R score compared to Caucasian patients 28. The importance of IPSS-R 22 is indubitable to define overall survival and risk of AML transformation in MDS, but it is very important to emphasize the unique profile of each population, primarily due to the many different population heterogeneity and miscegenation. It is expected the IPSS-R updates will consider the distinct genetic characteristics of each population, which would be important for definition of risk profile and the development of personalized medicine concepts. Our report reinforces that ethnic differences and regional influences may play a role in the pathogenesis and prognosis of MDS.
This study presents limitations. The number of patients evaluated in each country may not be truly representative of Brazil or Argentina but due to the low economic power of each nation, the researchers detected very important difficulties to collect DNA samples and perform genotyping by TaqMan methodology. This must be pointed out. Brazil and Argentina researchers do not have access to grants routinely and this compromises our evaluations, what creates difficulty obtaining higher number of cases than here presented.
The present study reaffirms MDS is a heterogeneous disease, showing the presence of polymorphic variants of XPA, XPC, XPD, and XPF genes with differences between Brazilian and Argentinean populations, suggesting these polymorphisms may help to understand the heterogeneity of MDS, creating possible new prognostic markers. In this study, we only evaluated nucleotide excision repair pathway enzymes (NER) which are an essential mechanism for single-stranded DNA repair. These results here presented open new possibilities to study functional polymorphisms of genes related to double strand breaks such as homologous recombination, single strand annealing and non-homologous end-join in different countries.
This work was supported by CAPES, CNPq (PRONEX) and FUNCAP.