Hemoglobin is an essential biological component of human physiology and its production in red blood cells relies upon proper biosynthesis of heme and globin protein. Disruption in the synthesis of these precursors accounts for a number of human blood disorders found in patients. Mutations in genes encoding heme biosynthesis enzymes are associated with a broad class of metabolic disorders called porphyrias. In particular, one subtype – erythropoietic protoporphyria – is caused by the accumulation of protoporphyrin IX. Erythropoietic protoporphyria patients suffer from photosensitivity and a higher risk of liver failure, which is the principle cause of morbidity and mortality. Approximately 90% of these patients carry loss-of-function mutations in the enzyme ferrochelatase (FECH), while 5% of cases are associated with activating mutations in the C-terminus of ALAS2. Recent work has begun to uncover novel mechanisms of heme regulation that may account for the remaining 5% of cases with previously unknown genetic basis. One erythropoietic protoporphyria family has been identified with inherited mutations in the AAA+ protease ClpXP that regulates ALAS activity. In this review article, recent findings on the role of ClpXP as both an activating unfoldase and degrading protease and its impact on heme synthesis will be discussed. This review will also highlight the role of ClpX dysfunction in erythropoietic protoporphyria.
Heme is a chemical moiety that is essential for all life.1–6 In humans, heme is most commonly known for its role in red blood cells (RBCs) as the oxygen-binding prosthetic group in hemoglobin. Given its importance in RBC physiology, defective heme metabolism is strongly associated with hematologic diseases in humans.
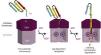
With few exceptions, heme is synthesized via a highly conserved set of eight enzymatic reactions. The first committed step of heme biosynthesis involves the condensation of glycine and succinyl-CoA in the mitochondria to generate δ-aminolevulinic acid (ALA) utilizing the enzyme δ-aminolevulinic acid synthesis (ALAS1). ALA is then transferred into the cytosol where it is converted to tetrapyrrole coporphyrinogen III (CPgenIII) by four enzymes – ALA dehydratase (ALAD), hydroxymethylbilane (HMBS), uroporphyrinogen III synthase (UROS), and uroporphyrinogen decarboxylase (UROD).7 After entering the mitochondrial inner membrane space, the enzyme coproporphyrinogen oxidase (CPOX) catalyzes the conversion of CPgenIII to protoporphyrinogen IX (PPgenIX).8 PPgenIX is then transported into the mitochondrial matrix where it is the reactant for protoporphyrinogen oxidase (PPOX), to produce protoporphyrin IX (PPIX). The penultimate enzyme in the cascade, ferrochelatase (FECH), is responsible for the final insertion of iron into PPIX to form heme (Figure 1).9
Highly conserved heme/protoporphyrin IX (PPIX) synthesis in eukaryotic cells. Precursors, glycine and succinyl-CoA, undergo 8 enzymatic reactions with δ-aminolevulinic acid synthesis (ALAS), ALA dehydratase (ALAD), hydroxymethylbilane (HMBS), uroporphyrinogen III synthase (UROS), uroporphyrinogen decarboxylase (UROD), coproporphyrinogen oxidase (CPOX), protoporphyrinogen oxidase (PPIOX), and ferrochelatase (FECH) in both the mitochondria and cytoplasm to form heme.
Figure illustration courtesy of Johannes G. Wittig (Technische-Universität-Dresden, Germany). Reprinted with permission from Yien et al.37 and Oncotarget (Impact Journals, LLC).
Erythroid heme production must meet the vast production of hemoglobin and accounts for roughly 85% of the daily heme synthesis.2 All heme biosynthesis genes are upregulated during RBC differentiation.2 However, RBCs also have unique mechanisms in addition to the conical heme biosynthetic pathway. For example, the first step of heme synthesis, involves an erythroid-specific isoform of ALAS (ALAS2) that becomes transcriptionally induced during RBC maturation. ALAS2 differs from ALAS1 through an iron responsive element (IRE) in ALAS2 that can interact with iron responsive proteins (IRPs) allowing for the association of ALAS2 to iron availability.10
The final enzyme, FECH, is also regulated in a red cell specific manner. FECH is an iron-sulfur cluster protein located on the inner mitochondrial membrane. In RBCs, FECH is post-translationally stabilized through the formation of a complex with mitoferrin1 and ABCB10.11–13 Recent work has also found that erythropoietin (EPO) signaling together with the GATA-1 transcriptional program regulates the phosphorylation and activation of FECH in RBCs.10–12 Based on the number of regulatory pathways converging on FECH in red cells, it is not surprising that FECH is a rate-limiting enzyme in erythroid heme biosynthesis.10
PorphyriasThe intermediates produced by each enzyme in heme synthesis are cytotoxic and their accumulation can have deleterious effects. Clinically, the accumulation of heme intermediates cause porphyrias – a diverse group of metabolic disorders. Patients with porphyria suffer from a variety of symptoms including cutaneous photosensitivity, behavioral changes, restlessness, insomnia, seizures, abnormal liver function, and several other life threatening concerns.2,14 The complex genotype-phenotype correlations in combination with environmental factors have given rise to a broad range of disease presentation. Mutations in nine genes are currently known to be associated with porphyrias. Of the nine, defects in seven genes are associated with hepatic porphyrias, such as porphyria cutanea tarda (PCT), and characterized by the accumulation of heme precursors in the liver.2 In contrast, defects in the remaining two genes – ALAS2 and FECH – encode enzymes particularly important in RBC biology and give rise to erythropoietic porphyrias with the distinctive feature of accumulating heme precursors in the bone marrow.2
FECH is the gene most commonly associated with erythropoietic protoporphyria (EPP), accounting for 90% of diagnosed cases.2 These patients have reduced FECH activity.2ALAS2-associated EPP mutations are activating and result in increased ALA production. Ultimately, both FECH and ALAS2 mutations lead to an accumulation of photoreactive PPIX. In EPP patients, accumulated PPIX reacts with sunlight resulting in the production of cytotoxic reactive oxygen species (ROS).4,15 For some patients, even a small amount of light exposure can trigger skin lesions, while severe sun exposure can result in edema, erythema, and wax-like scarring. Increased amounts of free PPIX can also cause patients to suffer from more life threating concerns such as gallstones, liver disease, and liver failure.14
Genetic basis of EPPEPP mutations can be inherited in an autosomal dominant, recessive, or X-linked fashion.2 The genetics of EPP have been extensively studied. EPP cases with FECH mutations most commonly exhibit an autosomal dominant pattern of inheritance in which the hypomorphic IVS3-48C FECH allele is found in all affected family members.16–18 When the other FECH allele is mutated, FECH enzymatic activity is substantially reduced to 15–25% of its normal activity level.2,17,19 Over 60 different FECH mutations including partial deletions, frame shifts, and substitutions, have been identified to be associated with the IVS3-48C allele and found to cause decreased FECH activity.16–18 The most common mutations found in this region associated with EPP involve partial deletions. Partial deletions within the FECH allele, varying among patients, have been found to alter the structure, activation regions, or stability of FECH resulting in reduced enzyme acitivity.17 Patients suffering from frame shift mutations result in either a defective protein or a premature stop codon.18 These changes disrupt the secondary structure of FECH, leading to a decrease in the effectiveness of the enzyme.18 Research preformed by Schneider-Yin et al. also found that FECH substitution mutations also cause EPP through changes of the structural components of the [2Fe-2S] cluster of FECH.16 The changes in the [2Fe-2S] cluster, found to play a structural role in FECH, destabilize the FECH enzyme structure resulting in a reduced level of enzymatic activity.16 Even though the mechanisms through which FECH activity is disrupted are diverse, they all hinder heme production and result in high PPIX.
Although the majority of EPP cases arising from FECH mutations are inherited in an autosomal dominant fashion, there are also a few FECH-associated EPP cases that present as autosomal recessive.20 In one family, two healthy parents, each containing a different mutation to the FECH allele, developed a minor increase in PPIX levels, but no other symptoms.20 When their children inherited both mutant alleles, both children developed severe EPP and presented with liver failure and extreme photosensitivity.20 In this particular case, the two mutations were a substitution and an insertion that reduced FECH activity to 57% and 64%, respectively.20 Although each mutation alone only partially hindered FECH activity, in combination, the mutations led to an extreme depletion of FECH activity, 13–25%.20 Even though these double mutations are rare, they can result in severe EPP.
Another 5% of EPP cases result from gain-of-function mutations in ALAS2, which presents as an X-linked disorder.2 These mutations cluster in the inhibitory C-terminal region of ALAS2 and cause either the elongation or deletion of the C-terminus on ALAS2.2,21 Alterations to the C-terminus of ALAS2 triggers increased ALAS activity resulting in abnormally high PPIX levels.21,22 However, the remaining 5% of EPP patients do not harbor FECH or ALAS2 mutations, suggesting additional genetic modifiers still exist.
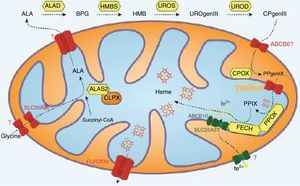
ClpX – a novel regulator of heme productionClpX is a mitochondrial AAA+ ATPase found in tandem with the mitochondrial tetradecameric peptidase hexamer, ClpP, to form the AAA+ protease ClpXP.23 Through in vitro studies using the model substrate casein, a model for ClpXP function has been developed.24–26 ClpX is involved in substrate binding, unfolding, and translocation to ClpP, which mediates protein degradation (Figure 2).23 Independent of ClpP, ClpX is still able to unfold proteins without degradation.23 While substrate recognition by ClpX requires only ATP binding, ATP hydrolysis is required for the unfoldase and translocase activities of ClpX.23 Although the biochemical function of ClpXP has been known for some time, physiologic substrates of ClpXP in mammalian cells are largely unknown.
ClpXP protease model of substrate recognition, unfolding, and degradation function. The peptide tag binds to the axial pore of ClpX. Subsequently, ClpX unfolds the substrate, with energy provided by ATP, and translocates the unfolded polypeptide to ClpP, which mediates proteolysis.
Reprinted with permission from Baker et al.23 and Biochim. Biophys. Acta (Elsevier, Inc.).
This has recently changed with a series of elegant studies using both yeast (S. cerevisiae) as well as vertebrate models.27 Yeasts lack ClpP and, thus, they are an excellent system to study ClpX unfoldase function without substrate degradation. Researchers, first, found that the yeast equivalent of ClpX, MCX1, promoted the activity of HEM1, which is the yeast homolog of ALAS1.23,27 Similarly, Kardon et al. discovered that mammalian ClpX, independent of ClpP, activated ALAS2 in vitro by facilitating PLP insertion. Based on these results, the authors proposed that ClpX activates ALAS2 in mammalian cells. In support, expression of ClpX but not ClpP was induced in maturing murine erythroid cells.27 Introduction of mammalian ClpP led to a decrease in ALAS activity, suggesting that active ALAS2 is a substrate for ClpXP degradation.27 Together these findings led the authors to propose that ClpXP plays a dual role in heme biosynthesis with ClpX as an activator of ALAS function and ClpP as a repressor. To further confirm ClpX as an activator, Kardon et al. showed that ClpX was required for efficient erythroid heme synthesis in zebrafish (Danio rerio) embryos. Morpholino knockdown of the zebrafish homolog, clpxa, in zebrafish embryos (termed morphants) resulted in a substantial decrease in hemoglobinization and the number of erythroid cells in vivo.25 The anemia found in ClpX morphants was rescued by ALA supplementation that by-passes the block in ALAS activity. These results suggest that the activation of ALAS by ClpX is a conserved pathway and represents an important step in heme synthesis.25
Further work by Kubota et al. has shed light into how ClpXP functions in both capacities and when it is important to do so.25 Using the human HepG2 liver cell line, the authors found that siRNA-mediated knockdown of ClpX led to a robust accumulation of ALAS1. This is consistent with the idea that ClpX is required not only for substrate binding and unfolding, but also shuttling to ClpP for degradation.25 Interestingly, similar inhibition of ClpP expression with siRNA did not result in ALAS1 accumulation. ALAS1 levels were only increased following prolonged knockdown of ClpP (ten days). This raises the possibility that additional proteases may be responsible for ClpX-dependent turnover of ALAS1. Alternatively, the mechanisms through which ClpXP regulates ALAS1 may differ from ALAS2.23,25,27 It is also important to note that the authors did not measure ALAS1 activity in ClpP-deficient cells.25 There is evidence to indicate that ClpP inhibits the ATPase activity of ClpX, which is crucial for ClpX unfoldase function.25,27 Presumably, in this fashion, ClpP would inhibit ALAS1 activity by blocking PLP insertion without affecting steady state ALAS1 expression. Nevertheless, based on these findings, Kubota et al. propose that ClpXP constitutes part of the negative feedback loop in which excess heme triggers ALAS1 degradation.
ClpX is a novel EPP geneRecently, a family was identified with a history of EPP but no mutations in FECH or ALAS2 that was inherited in an autosomal dominant fashion. Additional analysis found a common heterozygote ClpX missense mutation in all affected family members. This substitution of glycine with aspartic acid at position 298 of ClpX is located in the conserved Walker A sequence motif involved in ATP-binding.23 This ATP binding region is thought to be absolutely essential for the formation of the ClpXP complex.23,27 The alteration of an amino acid at the ATP binding region, according to models found by Baker et al. and Kardon et al., can cause an alteration in ClpX-ClpP interaction resulting in a variation in the translocation and degradation of both ALAS1 and ALAS2 producing altered heme levels.23,27
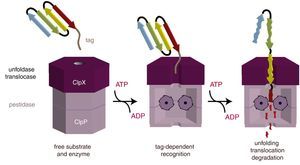
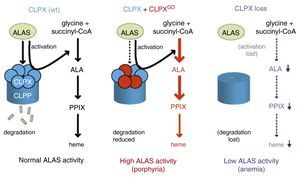
When the G298D ClpX mutant protein was exogenously overexpressed in murine erythroid cells, ALAS activity was increased with concomitantly higher PPIX levels.28 Interestingly, in vitro studies showed that the G298D mutation severely compromised ATPase activity of mutant ClpX. When mixed together in vitro with wild-type ClpX, mutant ClpX was also able to inhibit wild-type ATPase activity.28 Since ATP hydrolysis is required to form the ClpXP complex, the authors propose that mutant ClpX is unable to shuttle ALAS2 to ClpP for degradation; but retains the ability to unfold ALAS2 to allow PLP insertion (Figure 3). These results are significant for several reasons. First, the ability of mutant ClpX to inhibit wild-type function suggests that it is a dominant negative and provides a mechanistic explanation to the observed pattern of inheritance. Second, high ALAS activity even with diminished ATPase function suggests that ATP hydrolysis is not required for the unfoldase properties of ClpX or PLP insertion. It would be interesting to test whether this aspect of mutant ClpX also holds true with model substrates in vitro or if this is an effect specific for ALAS2. Furthermore, if ClpP is, indeed, a negative regulator, it is tempting to speculate that it too would be a candidate gene mutated in EPP. To date, the phenotype of ClpP knockdown or deletion remains unexplored.23,25,27,28 The potential role of ClpP in EPP and heme biosynthesis remains an exciting avenue of future research.
A model for how ClpXP regulates ALAS function and heme production. Under normal conditions (left), ClpX unfoldase activity is needed for PLP insertion and activation of ALAS proteins. ClpP mediated degradation constitutes negative feedback regulation in response to high heme levels.27 In EPP patients with the heterozygous G298D ClpX dominant negative mutation, active ALAS2 is unable to be shuttled to ClpP for degradation, resulting in excess production of intermediates (middle).27 In contrast, the complete absence of ClpX not only reduces degradation but also fails to activate ALAS. This causes a profound loss of heme production and anemia (right).26
Reprinted with permission from and Yien et al. and Proceedings of the National Academy of Sciences (PNAS).
Another remaining issue is whether ALAS2 gain-of-function mutations in EPP affect its interaction with ClpX. This is even more important since it has been proposed that ClpXP is a critical mediator of heme-dependent negative feedback regulation on ALAS1 activity.25 Additional studies are necessary to better understand the interplay between ClpX and mutant ALAS2. It is also tempting to speculate that the ClpX EPP mutations have disrupted heme feedback regulation. Nevertheless, ClpX mutations are now being recognized as a third variant of EPP. Whether ClpX mutations account for the entirety of the remaining cases of EPP with no known genetic cause is still unknown. Additional genetic studies are needed to determine the complete prevalence of ClpX mutations in EPP.
Current EPP treatmentsCurrent treatment options for patients are very limited. These options include avoiding sunlight exposure, reducing effect of sunlight exposure with oral β-carotene, administering porphyrin absorbents, such as cholestyramine and activated charcoal, transfusions, and in extreme cases liver or bone marrow transplant.2,29 Unfortunately, almost all current treatment options only alleviate the symptoms rather than addressing the pathogenesis of the disease. Only a bone marrow transplant has been found to cure a patient with EPP, but finding an efficient treatment plan and the stage to intervene has still not been fully determined.29
A proposed treatment for EPP involves the administration of ALADs to decrease intermediates upstream of PPIX. While this treatment has shown some promise in mice trials, ALADs has been shown to cause anemia if prescribed in excess.30 Therefore, this regimen must be closely monitored for adverse side effects.
Developing new treatments for EPP patientsFor the majority of EPP patients with defective FECH, research needs to focus on boosting residual FECH activity. Emerging evidence indicates that FECH in RBCs is activated by EPO signaling. EPO receptor signaling triggers activation of protein kinase A (PKA) leading to phosphorylation and activation of FECH.31 Since EPP patients with FECH mutations retain minor FECH activity, increasing the activity of FECH through EPO administration could help lower PPIX levels.2 However, researchers also found that if a mutation is altering the position of the FECH gene that is required for phosphorylation by PKA there is no increased activity.31 Therefore, this strategy may only be effective if the phosphorylation site is intact.
In addition to the treatment plans discussed earlier, EPP patients, without a FECH mutation, have found beneficial results when treated with iron supplements.27,28,32 Iron supplements help clear excess PPIX by driving the metalation reaction catalyzed by FECH.27,28,32 However, long-term effects of continual iron supplementation are still being investigated.32 In addition, treating with iron supplements has only been met with limited success. This is likely due to feedback inhibition of iron uptake via the bone morphogenetic protein (BMP) signaling pathway. Excess circulating iron leads to BMP-mediated stimulation of hepcidin production that, in turn, blocks cellular iron uptake into cells.33 Thus, although exogenous iron supplements place more iron in circulation, homeostatic mechanisms limit the extent of iron entering the cell. As a result, BMP inhibitors, such as inhibin and BMP-3, have been a major focus for research.34 Current efforts have failed to find a BMP inhibitor that is both potent and has a high enough selectivity to be effective in patients.35
Another approach is to use small molecules to increase iron uptake. In a recent small molecule screen, hinokitiol, found naturally in the leaves of a Japanese cypress tree, was found to bind and transport iron in and out of the cell.36 Hinokitiol has been shown to rescue zebrafish mutants with defects in cellular iron uptake as well as intracellular iron transport.36 Although its effectiveness has not yet been tested for treating EPP, this study displays the possibility of using iron carriers in hematologic diseases and paves the way for the discovery of other small molecules.
Conflicts of interestThe authors declare no conflicts of interest.
We would like to thank members of our lab for insightful discussions. We would like to dedicate this review to Dr. Barry H. Paw, who was a tremendous mentor and role model. This work was supported by the Diamond-Blackfan Anemia Foundation (BHP) and the National Institutes of Health Research (P01 HL032262, BHP and JC).
Barry Paw, MD, PhD, was a pediatric oncologist at the Dana-Farber/Boston Children's Cancer and Blood Disorders Center and principal investigator at Brigham and Women's Hospital, Harvard Medical School. Dr. Paw received his undergraduate degree in Biochemistry from University of California, Berkeley and his doctorate from University of California, Los Angeles School of Medicine. He had a highly distinguished career and was a recipient of several awards, including the William Randolph Hearst Young Investigator Award in 2002, the Basil O’Connor Scholar Award from the March of Dimes Birth Defects Foundation in 2004, the Young Investigator President's Award from the International BioIron Society in 2005, and was elected into the American Society for Clinical Investigation in 2008. Dr. Paw was a true pioneer in the field of heme and iron metabolism, combining his expertise of zebrafish genetics and biochemistry to study red cell development. His research has resulted in the identification of several novel genes involved in red cell development and disease. Dr. Paw will be remembered for his contributions to the scientific community. However, even more importantly, he will be forever cherished as a caring mentor, a good friend, and an impeccable person unmatched in his strength of character and love of science.