Chronic graft-versus-host disease (cGvHD) not only remains the main cause of late mortality after allogeneic hematopoietic cell transplant, but also has the capacity of causing severe organ impairment in those who survive. The Notch, a highly conserved ligand-receptor pathway, is involved in many immunological processes, including inflammatory and regulatory responses. Recently, mouse models have shown that the blockage of canonical Notch signaling prevents GvHD.
Objective and MethodDue to the lack of data on the Notch pathway in human chronic GvHD, we sought to study the expression of NOTCH components in primary samples of patients who received allo-HCT and presented active cGvHD or a long-term clinical tolerance to cGvHD.
ResultsOur results showed a significantly lower expression of NOTCH components in both groups that received allo-HCT, independently of their cGvHD status, when compared to healthy controls.
ConclusionMoreover, there were no differences in gene expression levels between the active cGvHD and clinically tolerant groups. To our knowledge, this is one of the first studies performed in human primary samples and our data indicate that much remains to be learned regarding NOTCH signaling as a new regulator of GvHD.
Subsequent to the allogeneic hematopoietic cell transplant (allo-HCT), activation of donor immune cells by host alloantigens can trigger both beneficial graft-versus-tumor effects, as well as adverse graft-versus-host disease (GvHD). The GvHD is the major complication of allo-HCT,1 which significantly limits the success of this curative procedure. Despite prophylaxis and treatment, the GvHD contributes substantially to morbidity and mortality, affecting 40 to 60% of allo-HCT recipients.2
The NOTCH signaling is a highly conserved ligand-receptor pathway, with pleiotropic functions, regulating embryogenesis, differentiation and adult tissue homeostasis, including multiple lineage decisions in hematopoiesis and function of innate and adaptive immune systems.3 For instance, dendritic cells depend on Notch signaling for their terminal differentiation and maintenance.4 The NOTCH also plays a role in orchestrating T helper (Th) cell differentiation, which includes its role in many arms of Th polarization.5 Naive T cells can differentiate into Th1, Th2 and Th17, depending on cytokines and also on NOTCH receptors present, such as DLL1 for the Th1 phenotype and JAG1 for the Th2 phenotype.5 The NOTCH signaling further affects the CD8+T cell differentiation,6 regulatory T (Treg) cell function and B-cell activation.7
More recently, we have learned that Notch is a central regulator of alloreactivity across different mouse models of allo-HCT.8 The genetic and pharmacological loss-of-function approaches in evaluating the in vivo effects of Notch signaling in T cells resulted in a dramatically decreased acute GvHD in different allo-HCT models, without causing global immunosuppression.9,10 A preclinical study, using humanized antibodies to block Notch signals in alloreactive T cells, showed that, despite Notch-blocked T cells having impaired production of multiple cytokines, they preserved their proliferation and expansion in vivo, including FoxP3+Treg cells and that they also maintained their antileukemic activity.11 Recently, Radojcic et al., using a mouse model of sclerodermatous cGVHD, showed that the blocking of Dll1/Dll4–mediated Notch signals provided maximum protection when used early after transplantation in a preventative fashion.12 Overall, mouse models showed that the blockage of canonical Notch signaling prevents GvHD, suggesting a tolerogenic effect.
Herein, we evaluated the gene expression of NOTCH signaling components in peripheral blood mononuclear cells (PBMC) of patients who received allo-HCT and developed cGvHD or a long-term clinical tolerance to cGvHD.
Material and methodsSubjectsSeventeen patients who underwent allo-HCT at our institution and 13 healthy controls were enrolled in our study during the year of 2016. The Ethics committee approved the study and informed consent was obtained from all patients. Patients with cGvHD were classified according to the 2014 NIH consensus.13 We divided patients into two groups: long-term clinically tolerant individuals (those who had never developed cGvHD or who had withdrawn from all immunosuppression therapy (IST) for at least 2 years and had no signs or symptoms of active cGvHD at the time of blood collection (Supplementary Table 1) and those who presented active cGvHD (with or without IST).
Sample collection and processingWe used the green top heparin blood tubes for the sample collections. After obtaining informed consent, peripheral blood samples from healthy donors and patients submitted to allo-HCT were collected at our outpatient unit, for convenience. Patient samples were preferably obtained on their exams collection and medical appointment day. The peripheral blood mononuclear cells (PBMCs) were isolated using the Fnicoll-Paque gradient centrifugation. We extracted RNA using the mirVAnaTMmiRNA isolation kit (Ambion/Life Technology, California), according to the manufacturer's instructions, followed by reverse transcription.
Quantitative RT-PCR (qPCR)Expression of mRNA was detected by qPCR and performed in an ABI 7500 Sequence Detector System (Thermo Fisher Scientific, Waltham, MA, USA) with SybrGreen PCR Master Mix (Thermo Fisher Scientific) and specific primers for NOTCH1 (FW: CTGCGAGCGGCCCTA; RV: CCGTTCTTGCAGTTGTTTCC), NOTCH2 (FW: TGCAAGCAATCACAAGCGTC; RV: GCCCTTCATCATCGACCCAG), JAG1 (FW: GCTTCCAACGACACACCTGA; RV: ATTTGCCTCCCGACTGACTC), JAG2 (FW: CGGCTATTACTGTGATTGCATCC; RV: ACACACTGGTACCCGTTCA), DLL1 (FW: GACCTCGCAACAGAAAACCC; RV: GTCACACACGAAGCGGTAGG), DLL4 (FW: ATGTACTTGTGATGAGGGCTG; RV: ATTCTTGCATGGGGAGTGGT) and HPRT1 (FW:GAACGTCTTGCTCGAGATGTGA; RV: TCCAGCAGGTCAGCAAAGAAT). The relative gene expression was calculated using the equation (Livak Methods 2001), 2ΔΔCT. A negative ‘No Template Control’ was included for each primer pair. The dissociation protocol was performed at the end of each run to check for non-specific amplification. Three replicas were run on the same plate for each sample.
Statistical analysisThe statistical analysis was performed using the Graph-prism 8.0 software (GraphPad Software, San Diego, CA) and SAS System for Windows version 9.4 (Statistical Analysis System, SAS Institute Inc, Cary, NC, USA). For comparisons, the Fisher's exact test, Mann-Whitney test and ANOVA, followed by the post-hoc Bonferroni, were used. The Kaplan-Meyer, followed by the log-rank test, was used for the overall survival (OS).
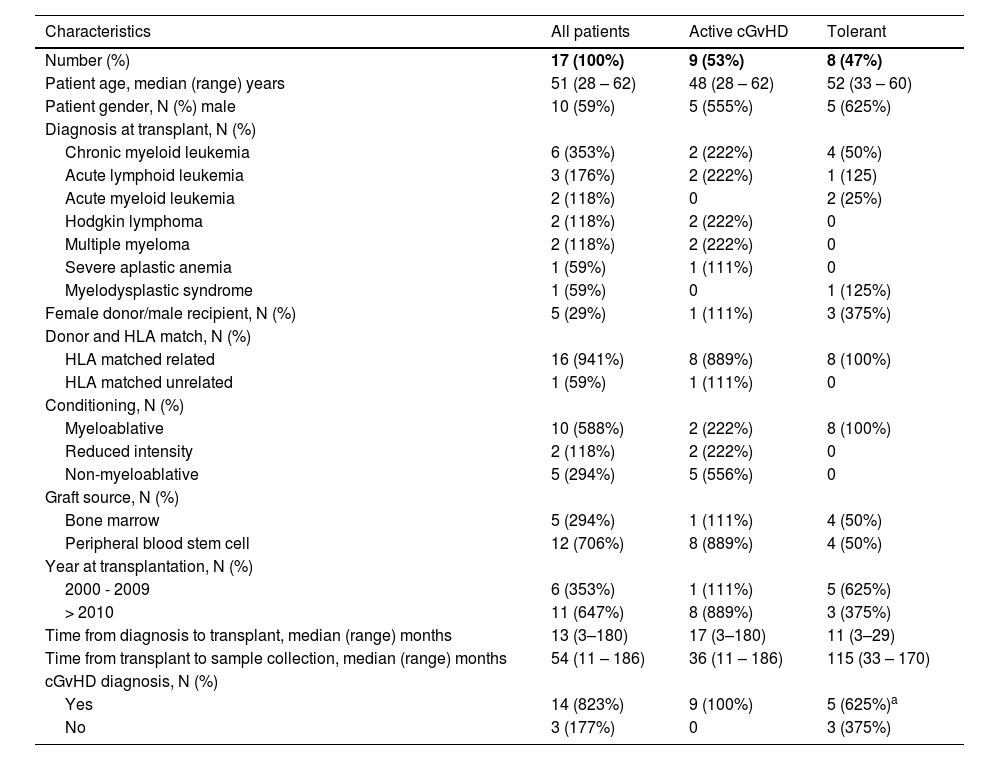
ResultsCharacteristics of the cohortAs presented in Table 1, 17 patients who underwent allo-HCT at our institution were enrolled; 9 (53%) had active cGvHD at the time of sample collection and 8 (47%) were clinically tolerant. The median time from the allo-HCT to the sample collection was 54 months (range: 11 - 186).
Patient characteristics.
| Characteristics | All patients | Active cGvHD | Tolerant |
|---|---|---|---|
| Number (%) | 17 (100%) | 9 (53%) | 8 (47%) |
| Patient age, median (range) years | 51 (28 – 62) | 48 (28 – 62) | 52 (33 – 60) |
| Patient gender, N (%) male | 10 (59%) | 5 (555%) | 5 (625%) |
| Diagnosis at transplant, N (%) | |||
| Chronic myeloid leukemia | 6 (353%) | 2 (222%) | 4 (50%) |
| Acute lymphoid leukemia | 3 (176%) | 2 (222%) | 1 (125) |
| Acute myeloid leukemia | 2 (118%) | 0 | 2 (25%) |
| Hodgkin lymphoma | 2 (118%) | 2 (222%) | 0 |
| Multiple myeloma | 2 (118%) | 2 (222%) | 0 |
| Severe aplastic anemia | 1 (59%) | 1 (111%) | 0 |
| Myelodysplastic syndrome | 1 (59%) | 0 | 1 (125%) |
| Female donor/male recipient, N (%) | 5 (29%) | 1 (111%) | 3 (375%) |
| Donor and HLA match, N (%) | |||
| HLA matched related | 16 (941%) | 8 (889%) | 8 (100%) |
| HLA matched unrelated | 1 (59%) | 1 (111%) | 0 |
| Conditioning, N (%) | |||
| Myeloablative | 10 (588%) | 2 (222%) | 8 (100%) |
| Reduced intensity | 2 (118%) | 2 (222%) | 0 |
| Non-myeloablative | 5 (294%) | 5 (556%) | 0 |
| Graft source, N (%) | |||
| Bone marrow | 5 (294%) | 1 (111%) | 4 (50%) |
| Peripheral blood stem cell | 12 (706%) | 8 (889%) | 4 (50%) |
| Year at transplantation, N (%) | |||
| 2000 - 2009 | 6 (353%) | 1 (111%) | 5 (625%) |
| > 2010 | 11 (647%) | 8 (889%) | 3 (375%) |
| Time from diagnosis to transplant, median (range) months | 13 (3–180) | 17 (3–180) | 11 (3–29) |
| Time from transplant to sample collection, median (range) months | 54 (11 – 186) | 36 (11 – 186) | 115 (33 – 170) |
| cGvHD diagnosis, N (%) | |||
| Yes | 14 (823%) | 9 (100%) | 5 (625%)a |
| No | 3 (177%) | 0 | 3 (375%) |
cGvHD: chronic graft-versus-host disease; w: with; w/o: without; HLA: human leukocyte antigen; N: number.
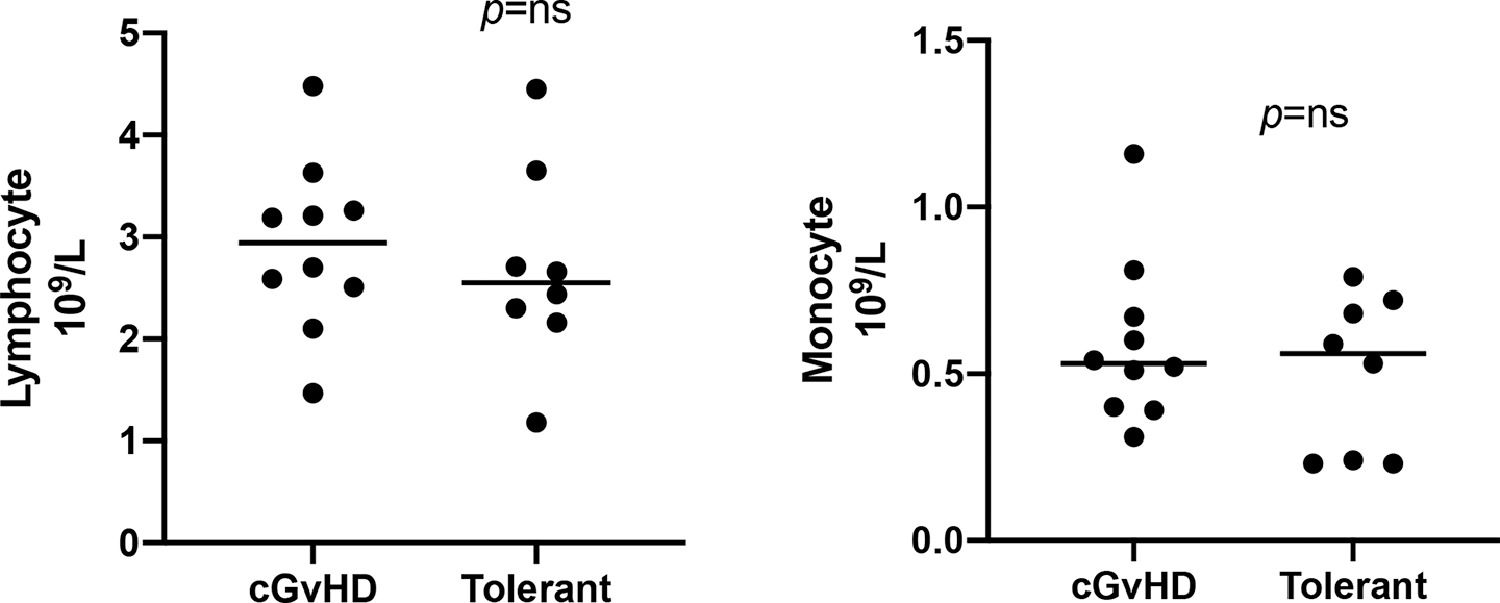
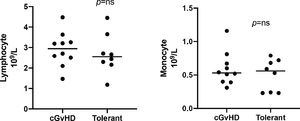
The median age was 51 years (range: 28 - 62) for the entire cohort. Ten (59%) were male and the most prevalent diagnosis was chronic myeloid leukemia (35%). Five (29%) of these patients were male with a female donor. Most patients had a matched related donor (94.1%) and had a myeloablative conditioning regimen (58.8%), with peripheral blood stem cells as the graft source (70.6%). Patients and transplant characteristics are shown in Table 1; the cGvHD characteristics at the time of the sample collection are shown in Table 2. The difference in lymphocyte counts and monocyte counts among the groups was not statistically significant at the time of the sample collection (Figure 3), and no patients showed any signs of infection at the time of the sample collection. Normal control samples were obtained from healthy blood donors and our service does not perform routine hemograms for this type of sample. Nevertheless, the lymphocyte and monocyte count in our cohort were within range, when compared to the Brazilian population, according to Rosenfeld et al.14
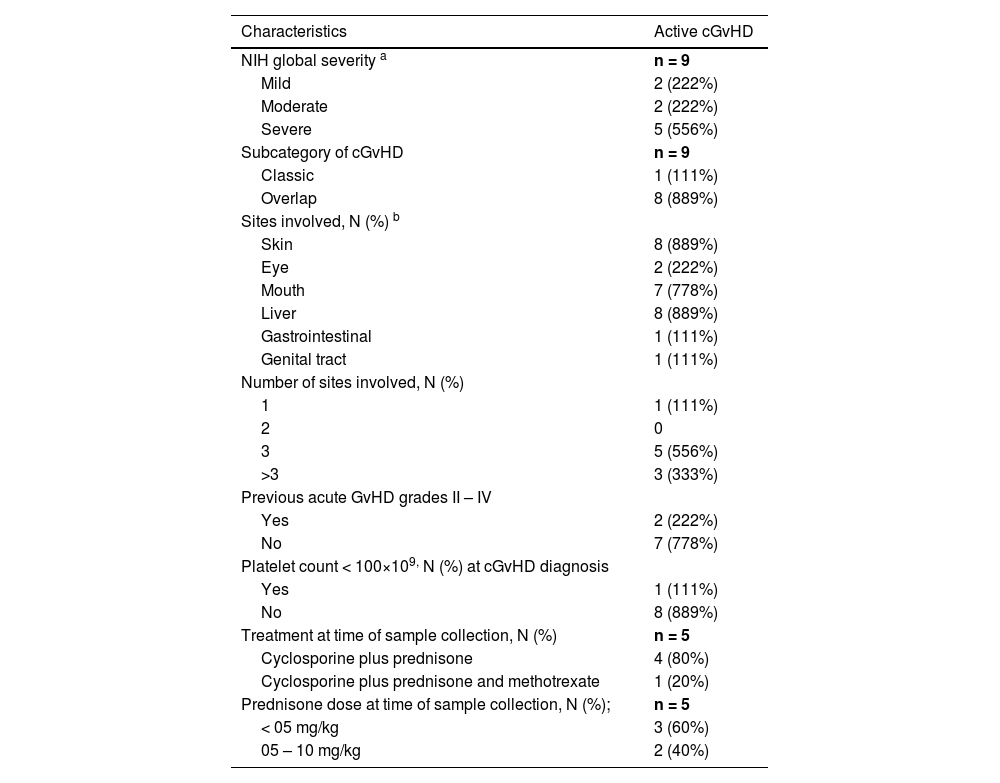
Chronic GvHD characteristics at enrollment.
| Characteristics | Active cGvHD |
|---|---|
| NIH global severity a | n = 9 |
| Mild | 2 (222%) |
| Moderate | 2 (222%) |
| Severe | 5 (556%) |
| Subcategory of cGvHD | n = 9 |
| Classic | 1 (111%) |
| Overlap | 8 (889%) |
| Sites involved, N (%) b | |
| Skin | 8 (889%) |
| Eye | 2 (222%) |
| Mouth | 7 (778%) |
| Liver | 8 (889%) |
| Gastrointestinal | 1 (111%) |
| Genital tract | 1 (111%) |
| Number of sites involved, N (%) | |
| 1 | 1 (111%) |
| 2 | 0 |
| 3 | 5 (556%) |
| >3 | 3 (333%) |
| Previous acute GvHD grades II – IV | |
| Yes | 2 (222%) |
| No | 7 (778%) |
| Platelet count < 100×109, N (%) at cGvHD diagnosis | |
| Yes | 1 (111%) |
| No | 8 (889%) |
| Treatment at time of sample collection, N (%) | n = 5 |
| Cyclosporine plus prednisone | 4 (80%) |
| Cyclosporine plus prednisone and methotrexate | 1 (20%) |
| Prednisone dose at time of sample collection, N (%); | n = 5 |
| < 05 mg/kg | 3 (60%) |
| 05 – 10 mg/kg | 2 (40%) |
cGvHD, chronic graft-versus-host disease; w: with; w/o: without.
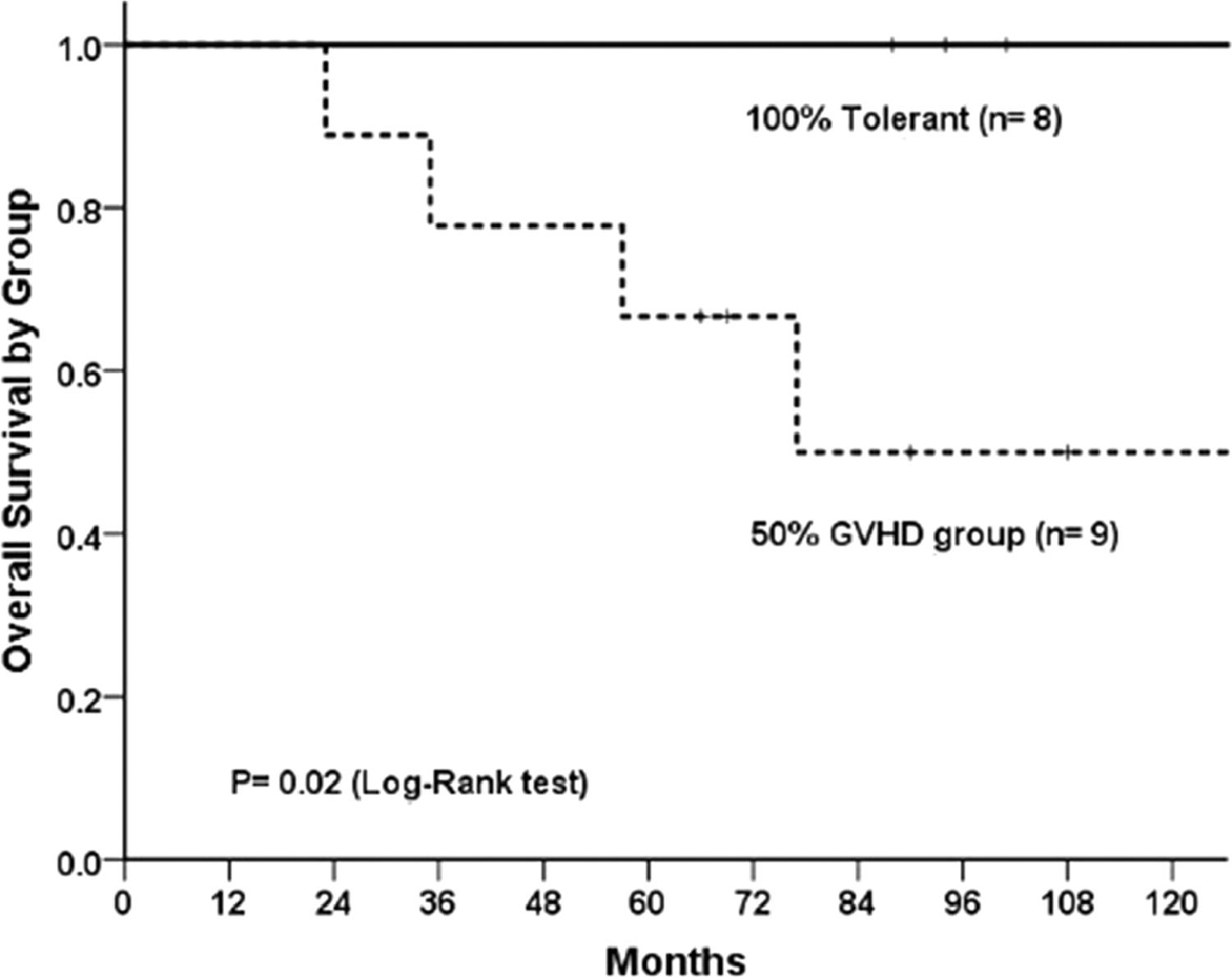
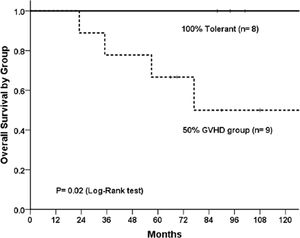
The overall survival with a 2-year follow-up was 100% for the clinical tolerant group and 50% for the cGvHD group (p = 0.02, Log-Rank test; Figure 1), which is in accordance with previously published outcomes.2,15 There was a difference in the conditioning-intensity distribution between groups (p = 0.005); a myeloablative regimen was applied for all clinical tolerant patients, whereas the cGvHD patients had an even distribution among the myeloablative, reduced intensity and non-myeloablative regimens.
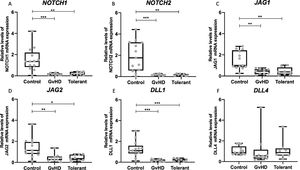
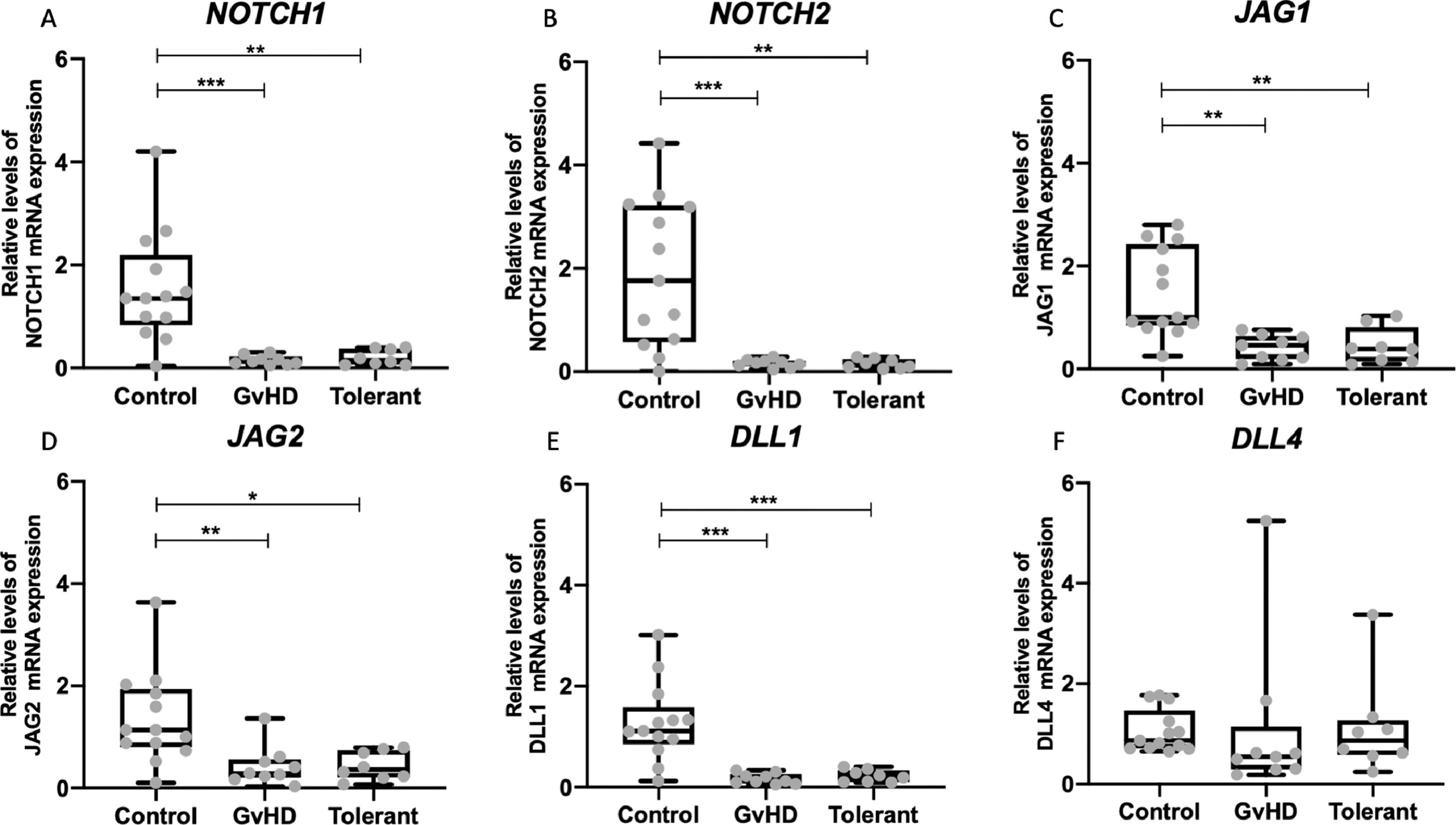
Lower expression of NOTCH components in PBMC of allo-HCT patientsThere was a significantly lower expression of NOTCH1, NOTCH2, JAG1, JAG2 and DLL1 in both groups that received allo-HCT, independently from their cGvHD status, when compared to healthy controls. Interestingly, there were no differences in gene expression levels between the active cGvHD and clinically tolerant groups (Figure 2).
Expression of NOTCH components in peripheral blood mononuclear cells in patients who received allo-HCT and control group Quantitative PCR was performed in cDNA from peripheral blood mononuclear cells for NOTCH1 (A), NOTCH2 (B), JAG1(C), JAG2 (D), DLL1 (E) and DLL4 (F) expression Relative expression was normalized by HPRT1 as the reference gene and results were analyzed using 2−ΔΔCT Horizontal lines represent medians The groups: control (n = 13), chronic graft-versus-host disease (cGvHD; n = 9) and tolerant (n = 8) are indicated *p < 005, **p < 001 and *** p < 0001; ANOVA test followed by Bonferroni post-test.
Herein, we analyzed for the first time, the expression of NOTCH components in primary PBMC of allo-HCT patients. The body of work that describes the role of Notch in alloimmunity is based on numerous experimental mice models, the majority having focused on the direct effects of Notch signaling on T cells. There are very few in vitro studies with human T cells 16,17; moreover, other cell types have emerged as critical elements in the GvHD setup, such as dendritic cells18 and, more recently, innate lymphoid cells (ILC).19 Recently, Poe et al. 20 evaluated, ex vivo, the B-cell receptor (BCR)/NOTCH2 synergy in B cells of patients, with and without cGvHD. Their results showed that the NOTCH2 blockade was able to eliminate the BCR activation, an important component of the cGvHD pathology.
Despite having observed a difference in the conditioning-intensity distribution between active cGvHD and clinical tolerant groups, these groups of patients had been enrolled in the study at least 11 months after the allo-HCT, which is more than enough time for the patients to reach total leukocyte and T cell chimerism, independent of the conditioning regimen intensity.21,22
There is compelling work indicating that inflammatory cytokines can induce the expression of NOTCH components, as well as the activation of the NOTCH pathway23 and it is well known that cGvHD is a complex immunopathological process that involves innate, as well as adaptive, immune cells.24 Therefore, the significantly lower expression of NOTCH components detected in both groups that received allo-HCT, when compared to healthy controls, as well as the similar downmodulation of NOTCH components in both groups of allo-HCT patients, was quite a surprise. Nevertheless, it is important to highlight that no mouse model of cGvHD developed as yet recapitulates the absoluteness of the characteristics that contribute to the pathology of cGvHD in humans: cytokine-mediated fibrosis, defective negative selection of T cells, deficiency of T reg cells, genetic polymorphisms and dysregulated B-cell responses.25
We are aware of the limitations of this study. We analyzed a small convenience sample of patients and we used whole PBMC, which contains lymphocytes ranging from 70 to 90%, but also contains other cell types.26
Our data indicates that much remains to be learned about NOTCH; for example, the exact mechanisms of NOTCH modulation in T cells have not yet been elucidated.27 Moreover, emerging roles for NOTCH signaling in innate and innate-like lineages, such as classical dendritic cells and ILC, are likewise coming into view.28 Animal models and in vitro studies pointed to Notch as a new major regulator of alloreactivity. However, our preliminary results showed that the expression of key components of NOTCH signaling is significantly downmodulated in the PBMC of patients who received the allo-HCT, when compared to the healthy group, along with no differences in gene expression levels between the active cGvHD and clinically tolerant groups.
ConclusionIn conclusion, our preliminary results showed that the expression of key components of the NOTCH signaling is downmodulated in the PBMC of patients who received the allo-HCT. However, until the NOTCH signaling is evaluated at the protein expression level in primary samples, whether or not the NOTCH signaling is critical for human allogeneic lymphocytes during the GVHD, remains unclear.
The authors would like to thank Dr. Nicola Conran and Raquel Susana Foglio for the English review and Eliana Miranda and Cleide Aparecida Moreira Silva for the statistical analysis. This study received financial support from the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP).