The myelodysplastic syndromes (MDS) are clonal stem-cell disorders characterized by ineffective haemopoiesis leading to blood cytopenias and by progression to acute myeloid leukemia (AML) in a third of patients.1 Diagnosis of MDS is based on examination of the blood and bone marrow (BM), which shows blood cytopenias and hypercellular marrow with dysplasia, with or without an excess of marrow blasts.2 The detection of chromosomal abnormalities in MDS supports the diagnosis, classification, prognostic stratification, therapy option, treatment monitoring and better understanding of the biology of the disease. The main chromosomal abnormalities of MDS are losses and gains of genetic material, and these changes are among the most important prognostic parameters in IPSS-R.3 As a consequence, major advances have been achieved in the treatment and survival of patients. However, nearly half of the patients present a normal karyotype, which prevents their further characterization.1 Therefore, there is the desire to increase the abnormalities detection rate by other methods.
The single nucleotide polymorphism (SNP) array method is a sensitive technology used to perform high-resolution genome-wide DNA copy number analysis and to detect the copy-neutral loss of heterozygosity (CN-LOH).4 The SNP array method uses millions of markers (probes comprised by 25-mer oligonucleotides) across the entire human genome. There are two types of probes: non-polymorphic probes, used only for assessing copy number, and polymorphic probes, used to assess genotypes. For copy number analysis, patient DNA labeled with a fluorochrome provides signal intensity that, when compared to a set of reference DNAs, indicates whether there is a gain or loss of genetic material. For genotype analysis, the SNP has a single base pair substitution (A, T, C, or G) of one nucleotide to another, following which the alleles are arbitrarily designated as A for one SNP and B for the other SNP and reveal which of the genotypes (for example, AA, BB or AB) is present along the entire chromosome.5
Previous studies have shown that the addition of the SNP array to karyotype (KT) increases by 28% the detection rate of cytogenetic abnormalities. To substantiate this idea, we describe two cases of MDS to which the SNP array added valuable information to the karyotype.6
MethodsPatientsPatients described in this report were accompanied at the Hospital São Paulo of the Federal University of São Paulo, São Paulo, Brazil. The Ethics Committee of the institution approved the study, and written informed consent was obtained (CAAE 00547512.5.0000.5505).
Metaphase cytogeneticsG banding karyotyping (KT) or metaphase cytogenetics was performed on BM samples at the time of initial diagnosis using standard methods and the karyotype was described according to the International System for Human Cytogenetic Nomenclature (ISCN) 2016.
Single nucleotide polymorphism arrayThe SNP array was performed on BM and buccal cells samples. Two hundred and fifty nanograms of DNA was extracted and digested, amplified, purified, fragmented, labeled and hybridized using the Affymetrix CytoScan® HD Array; CEL files were created using the GeneChip® System 3000 7G according to the manufacturer's instructions. The CEL files were analyzed using the Chromosome Analysis Suite v3.0 (ChAS) software. Regions of copy number variations (CNVs) larger than 1 Mb and CN-LOH larger than 10Mb, identified by the ChAS Software or detected by visual inspection, regardless of gene content, are denoted as true aberrations, with the exception of those known to be normal genomic variants (present in the Genomic Variants Database [http://projects.tcag.ca/variation]) or found in constitutional (buccal cells) SNP array analysis. Aberrations that were identified by the SNP array were described according to the ISCN 2016.
Cases reportCase 1: a 64yo male patient, classified according to the WHO 2016 as myelodysplastic syndromes with excess blasts (MDS-EB), bone marrow histology showed grade II fibrosis.
Karyotype: 46,XY,del(5)(q15q33),del(17)(p11.2)[16]/46,XY[4].
SNP array: arr[hg19] 5q21.1q35.3(99,749,120-180,719,789)x1; arr[hg19] (7)x1; arr[hg19] (8)x3; arr[hg19] 12p13.33p12.3(173,786-19,670,207)x1; arr[hg19] 12p12.1p11.22(25,285,957-28,222,696)x1; arr[hg19] 12q22q23.3(93,071,445-104,654,711)x1; arr[hg19] (16)x1; arr[hg19] 17p13.3p11.2(525-22,125,370)x1-2; arr[hg19] (Y)x1 (Figure 1).
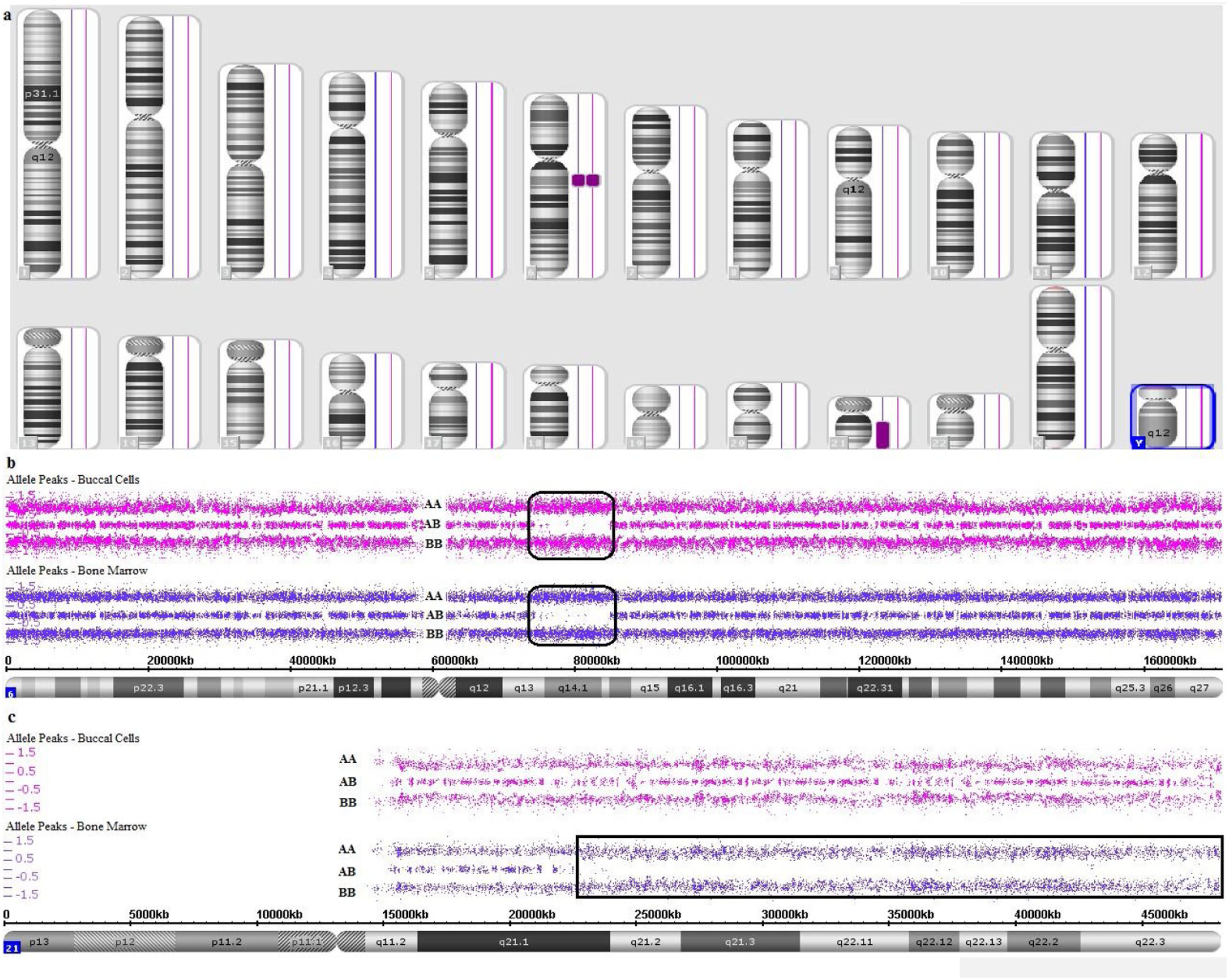
Genomic alterations detected by SNP array in case 1. a. Karyoview. b. Deletion in 5q21.1q35.3 (CN: 1.00). c. Deletion of one entire allele of chromosome 7 (CN: 1.00). d. Gain of one entire allele of chromosome 8 (CN: 3.00). e. Deletion in 12p13.33p12.3, 12p12.1p11.22 2 and 12q22q23.3 (CN: 1.00). f. Deletion of one entire allele of chromosome 16 (CN: 1.00). g. Mosaic loss in 17p13.3p11.2 (CN: 1.50). h. Loss of chromosome Y (CN: 0.00).
Case 2: a 74yo male patient, classified according to the WHO 2016 as myelodysplastic syndromes with excess blasts (MDS-EB), bone marrow histology showed grade II/lll fibrosis.
Karyotype: 46,XY[20].
SNP array: arr[hg19] 21q21.1q22.3(22,898,454-48,084,820) hmz (Figure 2).
Genomic alterations detected by SNP array in case 2. a. Karyoview of bone marrow (first line) and buccal cells (second line). b. Constitutional CN-LOH in 6q13q14.3 present in buccal cells and bone marrow SNP array analysis. c. Acquired CN-LOH in 21q21.1q22.3 present only in bone marrow SNP array analysis.
The SNP array detailed graphical results of all chromosomes (cases 1 and 2) can be seen in the Supplementary File.
DiscussionThe SNP array has the advantage of detecting genomic alterations regardless of the cell cycle, even when the cell is quiescent or has defective growth.7 It also enables the identification of CN-LOH (these regions are without heterozygosity, i.e. all the genetic variations within the regions have two identical alleles), submicroscopic amplifications and deletions that are not detected by KT.8 On the other hand, the SNP array does not allow for the identification of balanced translocations and polyploidy.6 In case 1, after SNP array analysis, some chromosomal abnormalities (−7, +8, −16 and −Y) were found in sporadic metaphases during the KT reanalysis, but had not initially been described because they did not meet the criterion to be considered as cytogenetic clones.9 The risk-stratification (IPSS-R) for this patient was intermediate, while addition of the SNP array results makes the risk-stratification very poor. Seven months after diagnosis the disease transformed into AML and the patient died. In case 2, the CN-LOH was detected by the SNP array in chromosomes 6 and 21. The CN-LOH in chromosome 6 was classified as constitutional since it was present in both BM and buccal cells, while the CN-LOH in chromosome 21 was classified as acquired, due to be present only in BM. The CN-LOH can be the mechanism responsible for homozygosity of mutations in critical genes located in the 21p region, such as the RUNX1 that encodes a protein that is a transcription factor critical in hematopoiesis. Indeed, sequencing of candidate genes in CN-LOH regions should be considered a priority in the search for driver mutations of MDS. Twenty-four months after diagnosis, the patient died due to other non-hematologic causes.
Whole genome scanning technologies have opened up a new road of investigation for chromosomal abnormalities in myeloid malignancies and also other neoplasms.10 The SNP array was first used as an investigative tool, which allowed for the identification of various common deleted regions and the discovery of several important gene mutations, exemplified by the CBL and TET2 mutations in MDS, and EZH2 mutations, in association with the CN-LOH of 7q.11 More recently, next generation sequencing (NGS) technology has also provided valuable information on molecular pathogenesis.2,12 As the cost of NGS tends to drop, molecular technologies will be increasingly applied in clinical and research labs. In this scenario, as a mature technology, with fully developed data analysis, such as the SNP array, it will play an important role in the clinical routine, especially in situations where diagnostic and prognostic represent a challenge. In Summary, the SNP array analysis may add value to KT non-informative results and occasionally reveal cryptic abnormalities not recognized by karyotyping.
FundingsThe SNP array test was supported by Grupo Fleury.
Conflicts of interestThe authors declare no conflicts of interest.